3 Lessons From Building With VectorAI DB at Hacklytics 2026

Summary
- An enterprise data marketplace is a centralized internal platform where teams can publish, discover, understand, and request access to data products.
- Its purpose is to break down data silos by making enterprise data visible, reusable, and easier to trust across departments.
- Typical features include cataloging, search, access workflows, quality indicators, and usage tracking.
- Main benefits include faster time to insight, less duplicate data work, stronger governance, better data quality, greater self-service, and improved collaboration.
- It helps organizations get more value from their data by turning scattered information into accessible, well-managed, business-ready data products.
A single wording change took a query in the FDA’s adverse event database (FAERS) from 1,532 results to just one. At Hacklytics 2026, Georgia Tech’s annual data science hackathon, the RxGuard team queried the FAERS database, containing over 31 million drug interaction reports, for Warfarin-Ibuprofen using four different phrasings: generic names, brand names, salt forms, and natural language. Each new query variant returned fewer results because the FAERS database could not understand language variation.
That finding was one of several that three teams surfaced while building with Actian VectorAI DB over a weekend. Each team picked a use case and tackled its retrieval, prediction, and generation constraints. Here are the healthcare problems they identified and the solutions they built.
Pharmacists querying the FAERS database for drug interactions miss critical safety signals because the system only recognizes exact keyword matches. RxGuard closes that gap with a semantic search engine that delivers investigation reports on dangerous interactions recorded across the database. The openFDA API pulls historical adverse event reports for 70 drug classes, all-MiniLM-L6-v2 embeds the reports as 384-dimensional dense vectors, and Actian VectorAI DB stores the embeddings. A single Vultr instance with Docker Compose powers the entire stack. When pharmacists enter a patient’s proposed medication and pre-existing condition, RxGuard retrieves the top 10 semantically similar FDA adverse event cases from VectorAI DB, along with severity risk scores and alternative medication recommendations.
Autoimmune diseases take four to seven years and four or more specialists to diagnose. Aura is a local-first clinical triage RAG engine. An XGBoost model, pretrained on public data from 100k patients, detects early signs of autoimmune disease from medical records with an AUC of 0.90. VectorAI DB serves as the local retrieval layer, storing PubMed records embedded using pritamdeka/S-PubMedBert-MS-MARCO, as 768-dimensional embeddings. An 8B parameter quantized model fine-tuned on general medical data and patient summaries produces clinical reports with citations from PubMed journals.
Most medical bills contain errors, but disputing one requires CPT code expertise, Medicare rate knowledge, and federal statute citations that many patients lack. PayBack is a multi-agent medical bill dispute system built on a LangGraph pipeline. Gemini 2.0 Flash extracts CPT codes from uploaded bills and benchmarks them against DoltHub’s transparent-in-pricing dataset, using either insurance-payer match or market-average fallback. A rules engine flags billing irregularities, while all-MiniLM-L6-v2 generates 384-dimensional embeddings for historical dispute cases locally. VectorAI DB stores these embeddings and pulls the three most relevant precedents as context for Gemini 2.5 Pro to draft the dispute letters.
Below, we share lessons from their development process, the critical search constraints they solved with VectorAI DB, and what their use cases reveal about vector databases in real deployment.
Keyword Search has Invisible Bias
For demos, keyword search gets the job done. But when real users query a domain-specific corpus using synonymous vocabularies, recall quality drops quickly. This is the vocabulary mismatch problem that keyword search exposes.
Domains like healthcare, finance, and law express the same terminology in multiple ways. The RxGuard team discovered this firsthand in the FAERS dataset: “The same drug appears under dozens of name variants, salt forms, foreign spellings, and typographical errors across 11.5 million reports. American researchers searching for ‘gastrointestinal hemorrhage’ get zero results because the correct MedDRA term uses British spelling.”
Keyword-based retrieval cannot recognize paraphrasing, topic similarity, or equivalence between expressions. As the RxGuard team puts it, “Keyword search doesn’t just return fewer results — it returns a systematically biased subset, and the bias is invisible to the researcher. Nobody knows what they’re missing because the missing cases never appear.” Support teams sifting through tickets, sales representatives searching product specs, and engineers digging through internal wikis all hit the same challenge. The fix is semantic matching.
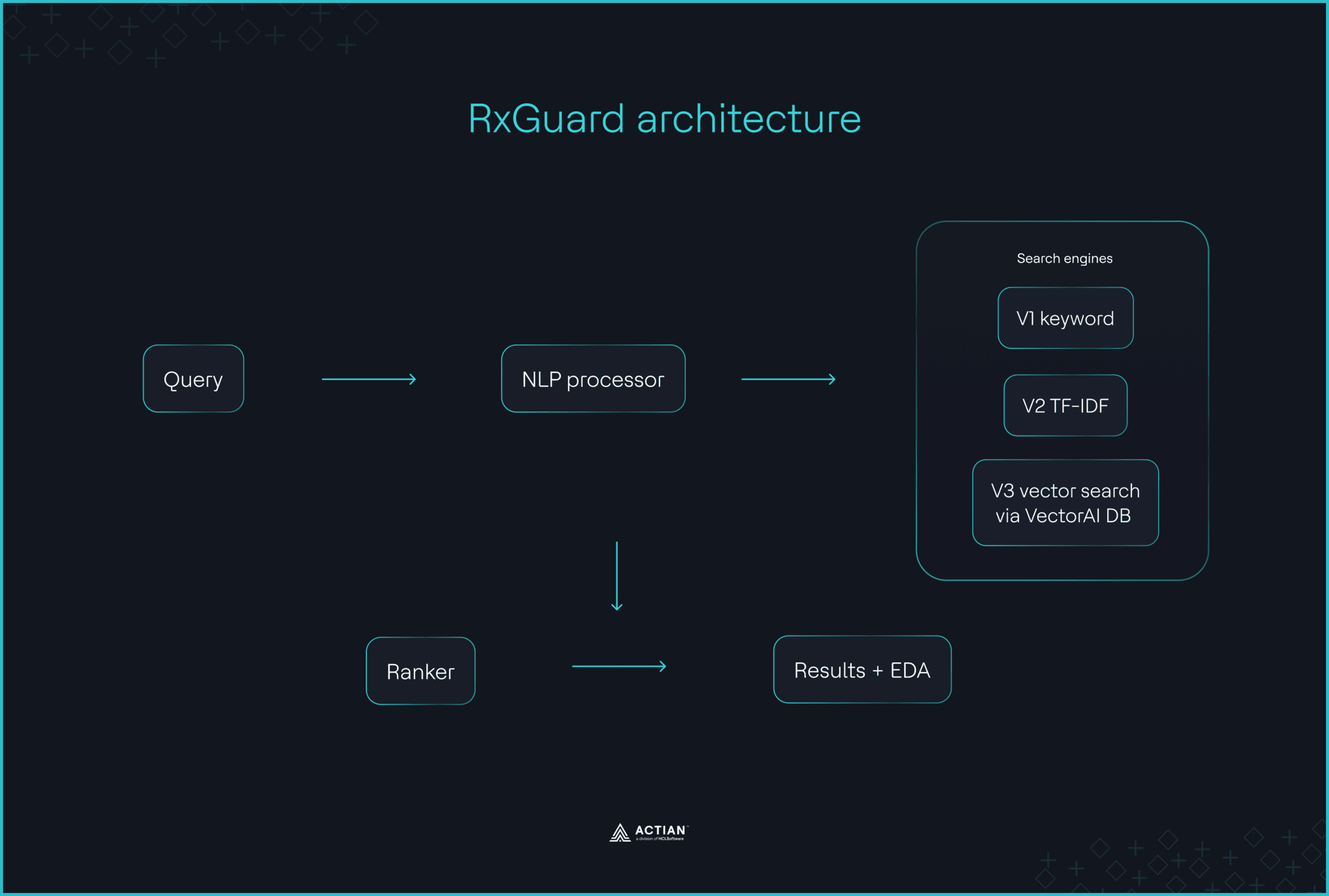
RxGuard used vector search via VectorAI DB as the primary retrieval mechanism, with optional pre-search severity filtering and drug-name matching applied as constraints, as shown below. Embeddings place synonyms and aliases close together in the vector space, so the system understands that “(…) ‘Coumadin,”warfarin,”‘and ‘blood thinner’ all mean the same thing.”
def search_faers(
query: str,
top_k: int = 10,
min_severity: int = 0,
drug_filter: str | None = None,
) -> list[dict]:
model = _get_model()
query_vector = model.encode(query).tolist()
f = Filter()
if min_severity > 0:
f = f.must(Field("severity_score").gte(min_severity))
fetch_k = top_k * 3 if drug_filter else top_k
with CortexClient(config.VECTORDB_ADDRESS) as client:
results = client.search(
config.VECTORDB_COLLECTION,
query=query_vector,
top_k=fetch_k,
filter=f if not f.is_empty() else None,
with_payload=True,
)
output = []
for r in results:
payload = r.payload or {}
# Post-filter by drug name if requested
if drug_filter:
drugs = payload.get("drugs", [])
if not any(drug_filter.lower() in d.lower() for d in drugs):
continue
output.append({
"rank": len(output) + 1,
"score": round(r.score, 4),
"doc_id": payload.get("doc_id", ""),
"text": payload.get("text", ""),
"drugs": payload.get("drugs", []),
"reactions": payload.get("reactions", []),
"severity_score": payload.get("severity_score", 0),
"patient_age": payload.get("patient_age", -1),
"patient_sex": payload.get("patient_sex", "unknown"),
})
if len(output) >= top_k:
break
return outputThis code block shows how the system transforms a natural language query into a ranked list of adverse event cases. VectorAI DB performs similarity search on the stored FAERS collection, retrieving the top 10 most relevant cases by default, regardless of the drug name phrasing. Metadata filters narrow results by severity score before retrieval, so only clinically relevant cases surface. For full code implementation, visit the GitHub repo.
Keyword search stops at matching explicit words, but retrieval efficiency depends on synonym awareness. And the solution isn’t foreign. As the RxGuard team puts it, “(…)the ingredients for this solution have existed for years: sentence transformers, vector databases, and the openFDA API. The problem wasn’t technical. It was that nobody had connected them to this specific use case.”
Vector Search Does One Job
The natural role of vector search in production systems is as a retrieval mechanism. Embedding models and approximate indexing methods make similarity search probabilistic. It surfaces conceptually similar documents, but cannot enforce specificity or interpret why those documents matter. In high-stakes industries where “close enough” is insufficient and explainability is non-negotiable, vector search alone should not serve as the decision layer.
The practical RAG and search pipeline for production deployment remains vector search for embedding retrieval, deterministic rules for decision-making, and an LLM for response generation. PayBack and Aura built this pipeline in distinct domains.
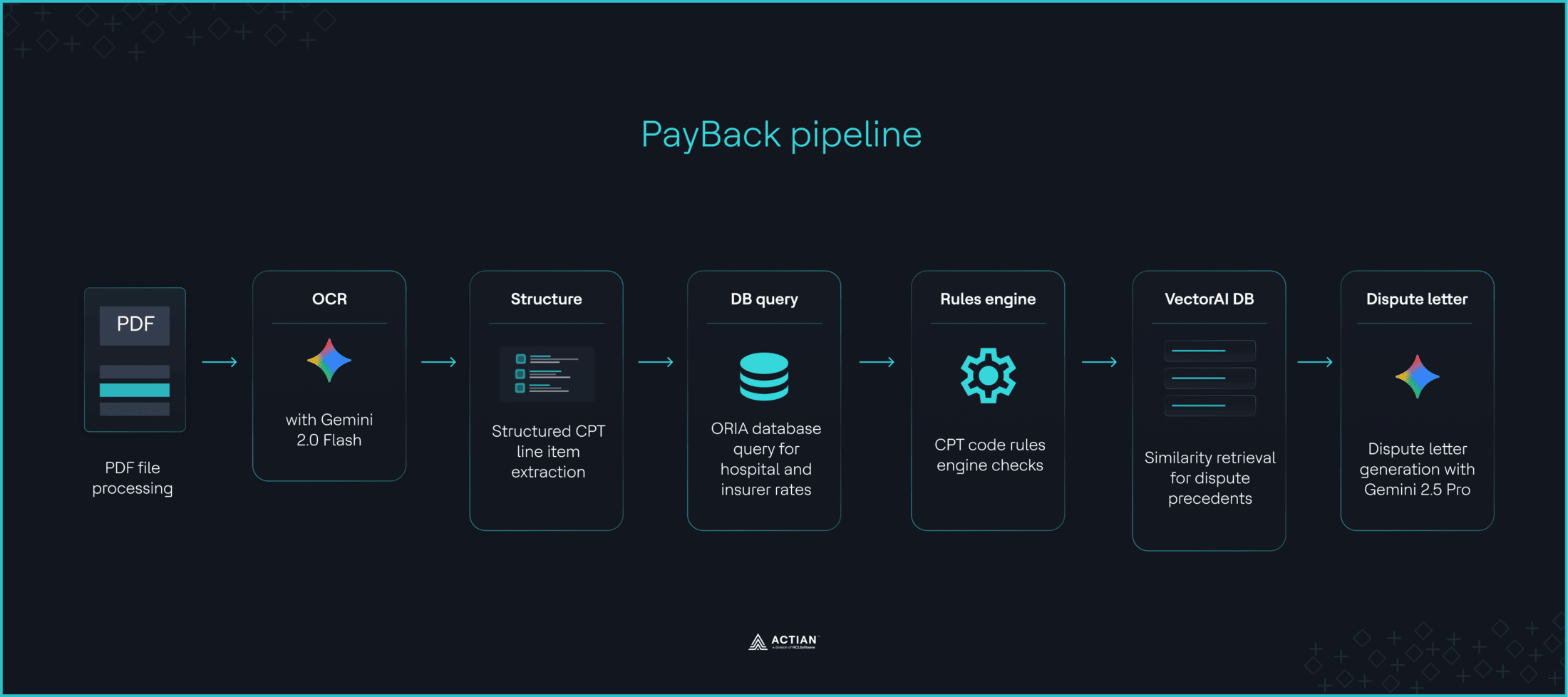
PayBack used a Python rules engine that flags “duplicates, quantity anomalies, extreme markup, facility fees, self-referral, and discharge-day billing.” VectorAI DB operated as a semantic retrieval and context enrichment layer that “fetches top‑3 similar historical dispute cases and injects them as context for Gemini prompts.” Gemini 2.5 Pro then “generated a formal dispute citing flags, DoltHub benchmarks, and precedent language.” This code snippet captures the handoff between PayBack’s rules engine and vector search. Visit the GitHub repo for the complete code.
if line_items:
try:
query = _build_precedent_query(line_items)
precedents = search_precedents(query, top_k=3)
similar_cases_context = _format_similar_cases(precedents)
print(f"[rules_engine] Fetched {len(precedents)} similar cases for context")
except PrecedentServiceError as exc:
print(f"[rules_engine] Precedent search unavailable, proceeding without: {exc}")The Aura team paired vector search with a hierarchical dual-scorer built on XGBoost. The model classifies structured patient features into disease clusters and identifies a likely condition within the predicted cluster. This classification embeds explainability into the system because Aura can trace which patient input drove the prediction and the confidence score.
VectorAI DB remained on the retrieval path, using patient feature context to surface the top five relevant PubMed literature that supports XGBoost’s output with clinical evidence. Here’s a snippet of how the Aura team configured VectorAI DB to store and filter PubMed chunks with metadata, rather than sweeping the full literature at query time.
with CortexClient(ACTIAN_HOST) as client:
client.create_collection(
name=COLLECTION_NAME,
dimension=EMBEDDING_DIM,
distance_metric=DistanceMetric.COSINE,
)
for batch_start in range(0, total, EMBED_BATCH):
batch_end = min(batch_start + EMBED_BATCH, total)
texts = rows["text"][batch_start:batch_end]
embeddings = model.encode(
texts,
batch_size=EMBED_BATCH,
normalize_embeddings=True,
).tolist()
for sub_start in range(0, len(texts), UPSERT_BATCH):
sub_end = min(sub_start + UPSERT_BATCH, len(texts))
i = batch_start + sub_start
client.batch_upsert(
COLLECTION_NAME,
ids=list(range(i, i + (sub_end - sub_start))),
vectors=embeddings[sub_start:sub_end],
payloads=[
{
"chunk_id": rows["chunk_id"][i + k],
"doi": rows["doi"][i + k],
"journal": rows["journal"][i + k],
"year": rows["year"][i + k],
"section": rows["section"][i + k],
"cluster_tag": rows["cluster_tag"][i + k],
"text": rows["text"][i + k],
"pmc_id": rows["pmc_id"][i + k],
}
for k in range(sub_end - sub_start)
],
)A translator model drafted the final patient summary and clinical SOAP note with DOI citations from the retrieved PubMed literature. Aura’s pipeline illustration below highlights how XGBoost produces the actual diagnosis prediction, and vector search adds justification. See complete code implementation on GitHub.
Use vector search to understand query intent, shortlist potential matches, and augment LLM responses. Add explicit logic when you need precision and output traceability. Collapse this separation, and search pipeline accuracy drops in production.
Sensitive Data has a Local Solution
Vector search belongs on-premises, where the data already lives. Regulations like HIPAA and GDPR impose strict requirements on where personally identifiable information (PII) and protected health information (PHI) must reside. For many industries, that means within the internal network.
Aura, PayBack, and RxGuard independently hit this data residency constraint. The Aura team struggled to source data during the hackathon because “Patient data is held very tightly by hospitals as per HIPAA regulations.” PayBack ran all-MiniLM-L6-v2 locally, keeping embeddings within the same boundary as the source documents. RxGuard plans a “Longer term, on-premises hospital deployment where HIPAA compliance requires that patient medication queries never leave the network, which is exactly the architecture Actian VectorAI DB was built for.”
Three teams converged on the same architectural decision. Vector search needs to run in a controlled environment from day one, and VectorAI DB keeps that architectural requirement lightweight. Aura spun up VectorAI DB locally with three commands:
git clone https://github.com/hackmamba-io/actian-vectorAI-db-beta
cd actian-vectorAI-db-beta
docker compose up -dThe team emphasized, “No patient data leaves the device. The vector database, model inference, and report generation all run on the user’s machine. This is non-negotiable for medical AI.”
RxGuard hosted NLP extraction, embedding generation, vector search, and DailyMed label retrieval on a single Vultr instance, with VectorAI DB running via Docker Compose. When compliance restricts sending private data to an external API or internal policies require inference to run locally, similar on-premises deployment logic makes the most sense.
A local Docker instance gives you full control over how sensitive data moves through ingestion and indexing to retrieval and response generation. Roles govern access to the vector store and every query leaves an audit trail. No confidential data crosses a network boundary without explicit authorization.
What These Projects Tell You About Your Next Build
Drug safety research, autoimmune disease detection, and medical bill disputes are three problems in one industry. Healthcare and life sciences sit on decades of structured clinical data, strict regulatory frameworks, and fragmented terminology. These three teams built working systems in a weekend. The use cases they picked are a fraction of what this industry needs.
Each project applied existing architecture for production-grade search systems. As the RxGuard team noted, “The ingredients for this solution have existed for years. The problem was that nobody had connected them to this specific use case.” Embedding models, vector search, deterministic rules engines, and local Docker deployment are not new. These teams recognized where each tool belonged and applied it to the right constraint.
The same problems they encountered exist in your domain. Vocabulary mismatch shows up in any corpus with inconsistent terminology, result traceability matters for systems that must justify every output, and data residency is a requirement in regulated industries.
If you are evaluating VectorAI DB for a domain-specific search tool or a RAG engine built around sensitive data, these three teams already validated the architecture you are considering. Test Actian VectorAI DB against your data using the local installation guide in the GitHub repo.
Join the Actian community on Discord to connect with other developers implementing vector search locally.





