How to Build a HIPAA Compliant AI Ecosystem Without the Cloud
Summary
- A BAA does not make cloud RAG automatically compliant, because the biggest privacy risks often come from the application layer, not just the provider’s infrastructure.
- The safer approach described here is to keep the entire RAG workflow on local hospital infrastructure, including ingestion, retrieval, generation, and logging.
- The system protects data by de-identifying content before ingestion, enforcing role-based access at query time, and preventing cross-department retrieval.
- It uses a local vector database and local models, so no patient data, embeddings, prompts, or logs leave the network.
- Local audit logging is essential because it creates a complete, verifiable record of who queried what, when, and under which role.
Healthcare cannot rely on cloud RAG because patient data leaves your network and your system logs, stores, and exposes it outside your control. You sign a Business Associate Agreement (BAA), connect your pipeline to a managed vector database, and assume compliance is complete. That assumption is wrong. The BAA covers the provider’s infrastructure. It does not cover what your application sends, logs, or exposes during retrieval and generation.
You remain responsible for every path your system sends Protected Health Information (PHI) through. A clinician query can leak sensitive data through logs. A system prompt can include patient context that your system stores outside your boundary. Weak access control allows retrieval results to expose records across departments. These risks exist in your application layer, not in the cloud provider’s scope.
American regulators now target this gap. In 2026, they flagged attack patterns like membership inference, where an adversary probes an AI system to confirm whether a patient’s data exists in the index. Cloud-hosted pipelines increase this risk because queries and embeddings move across external infrastructure. Audit requirements tighten further when logs live on third-party systems.
In this tutorial, you will build a clinical knowledge assistant that runs entirely on hospital infrastructure. It performs semantic search over clinical data, enforces role-based access at query time, and generates answers with clear citations. Every query stays inside your network, every access is logged locally, and no external API calls are required.
Why BAA is Not Enough
A BAA protects the cloud provider’s infrastructure, not how your system handles PHI during queries, retrieval, and generation. You remain responsible for every place PHI appears, moves, or gets stored inside your pipeline. There are multiple failure modes that make your system non-compliant, even when you sign a BAA.
Shared responsibility gap
The BAA stops at the infrastructure boundary. Your system controls what enters a prompt, what gets logged, and what leaves your network. If a clinician query includes PHI and your application logs it to an external service, you are responsible. If your retrieval step returns records across departments without strict filters, you have created an internal data breach. These failures happen in your code, not in the cloud provider’s scope.
For example, a physician searches “Show me similar cases to John Doe with early stage lung cancer.” Your application logs the full query to a cloud logging service for debugging. That log now contains PHI outside your network. The cloud provider did not leak it. Your application sent it.
Audit log ownership
HIPAA requires a complete audit trail for every access to PHI. When your vector database runs on third-party infrastructure, your system stores query logs and retrieval traces outside your control. You cannot guarantee completeness, retention, or isolation. Your security team cannot verify access patterns without relying on another provider’s system. That breaks your ability to enforce and prove compliance.
For example, your compliance team asks for a report of all oncology patient records in the past 30 days. Your vector database provider stores query logs on their platform with limited retention. Some logs are missing and others lack user-level metadata. You cannot produce a complete audit trail.
Membership inference exposure
Attackers can probe your system with targeted queries to determine whether a specific patient’s data exists in your index. This attack class is now a regulatory concern. Cloud-hosted indexes increase this risk because they expose a remote interface for repeated probing. A locally hosted index removes that external interface and limits access to your internal network.
For example, an attacker sends repeated queries like “Patients diagnosed with HIV in 2024 treated with drug X” and slightly modifies filters each time. They observe changes in response confidence and content. Over time, they infer whether a specific individual’s record exists in your dataset.
These failures show that a BAA does not ensure compliance. An on-premises deployment removes the third-party surface entirely and gives you full control over data flow, access, and auditability.
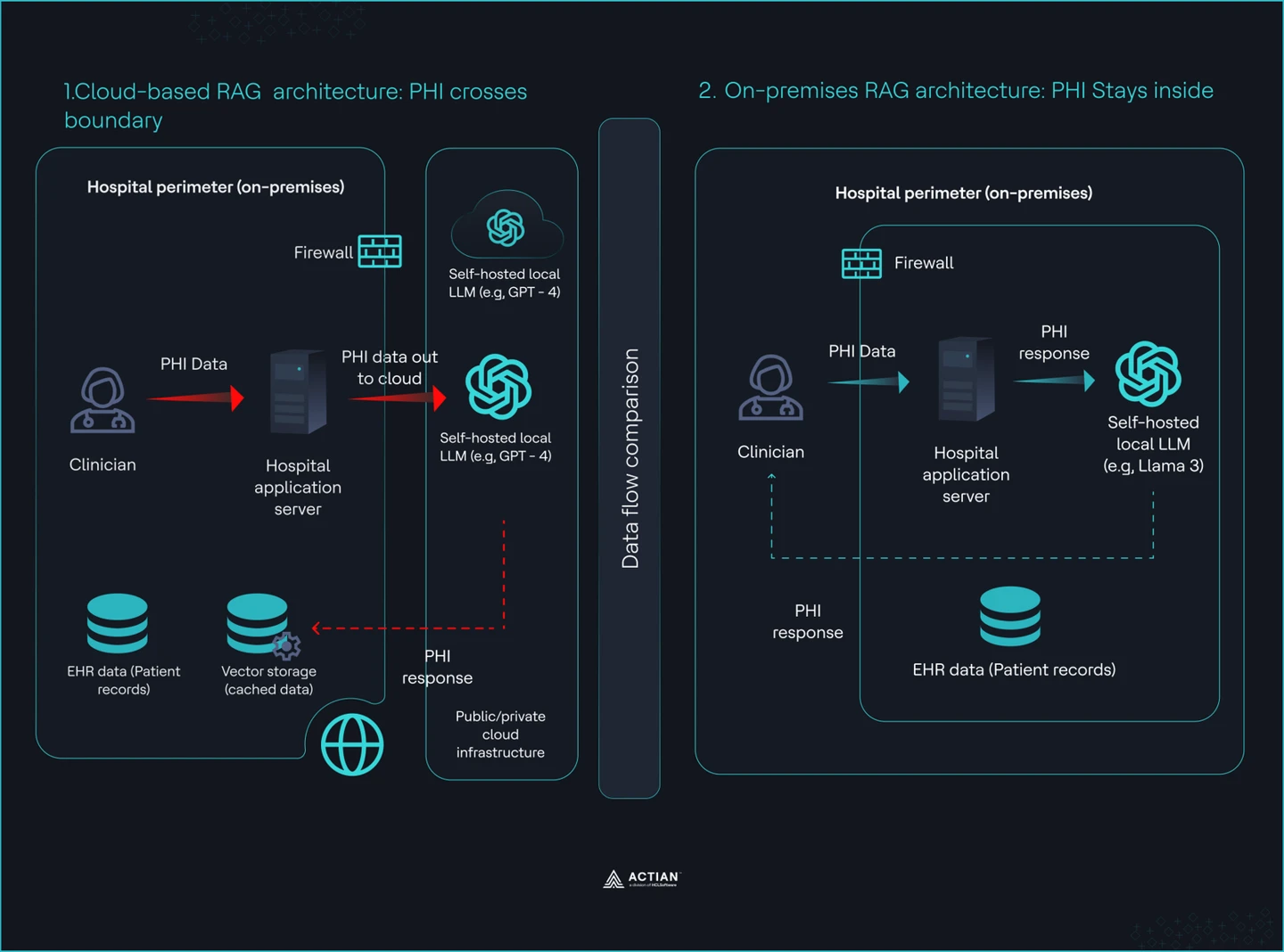
Split view showing Cloud vs. On-premises RAG architecture
What You are Building
In this section, you will build a RAG system with three layers:
Ingestion layer
You ingest clinical notes and treatment protocols into a controlled vector index with enforced data hygiene. You de-identify data before any processing. HIPAA Safe Harbor requires the removal of identifiers, while Expert Determination allows a statistical approach. You apply one of these before ingestion, not after. You then chunk documents into 512 token segments with 50 token overlap, generate embeddings using a local model, and store them in VectorAI DB with metadata.
You define a strict schema for every record. Each chunk includes document type, department, date, and author role. This metadata is not optional. It enables access control at query time and prevents cross-department leakage. Do not store raw documents without structure.
Query layer
You process clinician queries through a controlled retrieval pipeline. Every query passes through role-based access control before it reaches the index. A cardiology user can only retrieve cardiology data. A scheduling bot cannot access diagnosis notes. You enforce this with a MUST filter on department or patient cohort at the database level.
Run a hybrid search. Vector similarity retrieves semantically relevant chunks. Metadata filters restrict the result set. Pass the filtered context into a local LLM. The model generates an answer from retrieved data only and includes citations. Do not allow the model to invent or pull from external knowledge.
Audit layer
Log every interaction locally with full traceability. Each query writes a record that includes a timestamp, user ID, department, query text, and retrieved document references. This log lives on your infrastructure with defined retention and access policies. You do not rely on external logging systems.
You can reconstruct any access event from this log. You can answer who accessed what, when, and under which role. This satisfies audit requirements and gives your security team direct visibility into system behavior.
The entire system runs on commodity hardware inside the hospital network. The end-to-end architecture of the system is shown in the image:
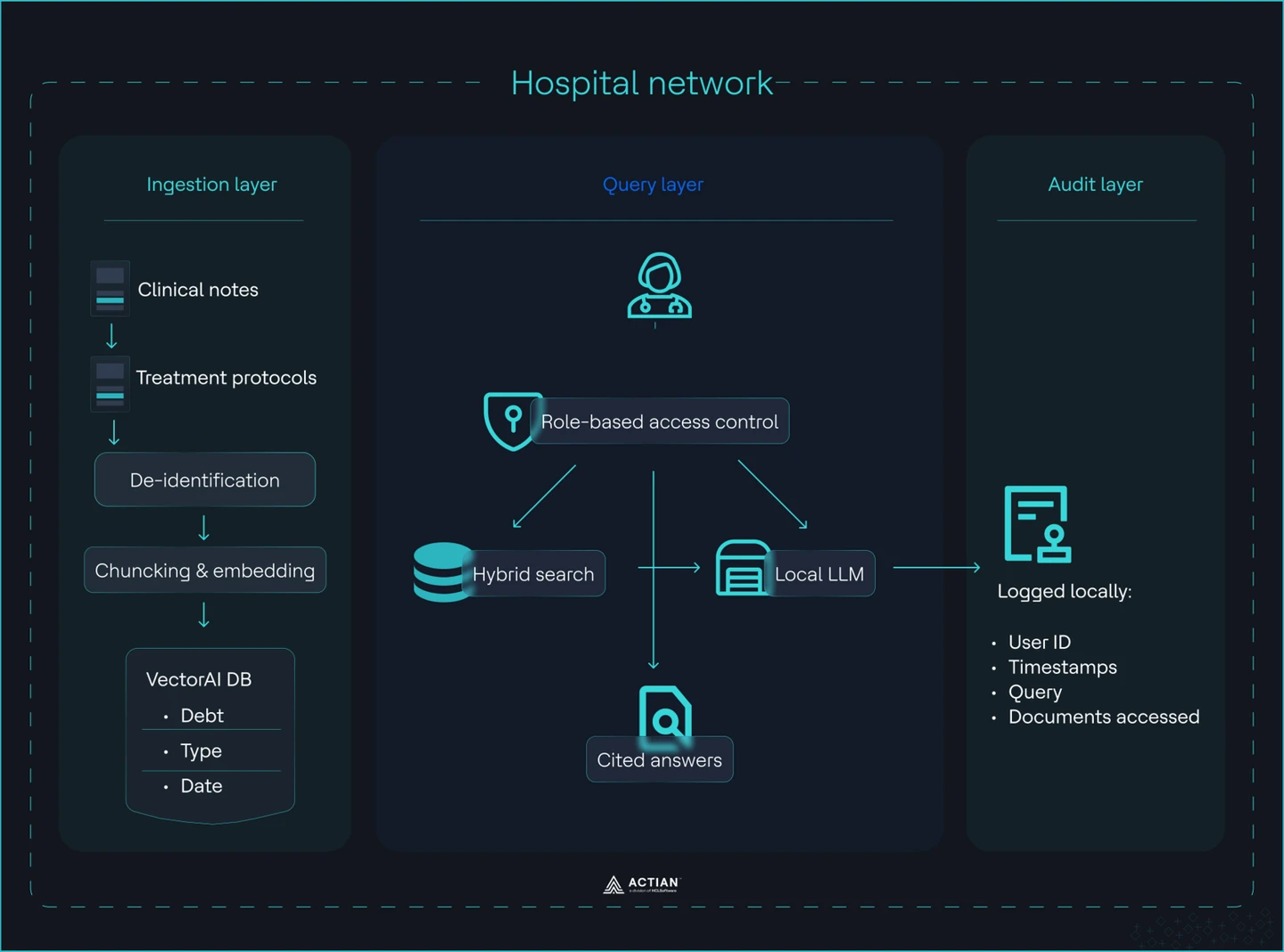
Hospital RAG system architecture
Building a HIPAA Compliant RAG Workflow
In this section, you will build a fully local RAG system that ingests clinical data, enforces access control, answers queries, and logs every interaction.
Prerequisites
To follow along, install the following tools on your local network:
Step 1: Deploy a vector database
You deploy a local instance of Actian VectorAI DB with persistent storage for both vector data and audit logs.
Create a docker-compose.yaml file:
services:
vectorai:
image: williamimoh/actian-vectorai-db:latest
platform: linux/amd64
container_name: vectorai_db
ports:
- "50052:50051"
volumes:
# vector data persists across restarts
- ./data:/app/data
# audit log lives on host — not inside the container
- ./audit_logs:/app/audit_logs
environment:
- VECTORAI_LOG_LEVEL=info
restart: unless-stopped
Run the service:
docker-compose up -dThe database starts and exposes port 50051 for local access. Vector data persists in ./data. Audit logs write to ./audit_logs on the host, which keeps all access records inside your network boundary.
Note:
- M3/M4 Apple Silicons might encounter a GRPC disconnection error without any container logs. In this case, disable Rosetta in Docker Desktop.
- VectorAI DB is under active development.
Step 2: Build the ingestion pipeline
Install the client library and run the ingestion pipeline to convert clinical documents into embeddings and store them in your local vector database.
Use uv for dependency management and execution. It is fast, reproducible, and avoids global Python state.
Download the Actian VectorAI client package. This creates a file actian_vectorai-0.1.0b2-py3-none-any
Initialize your project by:
uv init .
uv venvAfter initialization, install the Actian VectorAI package by:
uv pip3 install actian_vectorai-0.1.0b2-py3-none-anyAdd the embedding model dependency:
uv add sentence-transformersCreate a file ingest.py with the following contents:
import re
import hashlib
from actian_vectorai import VectorAIClient, Distance, VectorParams, PointStruct
from sentence_transformers import SentenceTransformer
# ── Config ────────────────────────────────────────────────────────────────────
VECTORAI_HOST = "localhost:50052"
COLLECTION = "clinical_docs"
EMBED_MODEL = "sentence-transformers/all-MiniLM-L6-v2" # 384-dim
VECTOR_DIM = 384
CHUNK_TOKENS = 512
OVERLAP_TOKENS = 50
# ── Synthetic clinical notes (replace with real de-identified corpus) ─────────
RAW_NOTES = [
{
"document_id": "card_note_001",
"document_type": "clinical_note",
"department": "cardiology",
"date": "2025-03-15",
"author_role": "attending_physician",
"text": """
Patient: [NAME REDACTED], DOB: [DATE REDACTED], MRN: [MRN REDACTED]
Chief Complaint: Chest pain radiating to left arm, onset 2 hours ago.
Assessment: Acute ST-elevation myocardial infarction confirmed on ECG.
History: Hypertension and type 2 diabetes. Started on aspirin 325 mg,
clopidogrel 600 mg loading dose, and heparin infusion per ACS protocol.
Plan: Emergency PCI. Beta-blocker therapy with metoprolol succinate
25 mg daily post-procedure. ACE inhibitor ramipril 5 mg daily initiated
24 hours post-PCI. Follow-up echocardiography in 6 weeks.
""",
},
{
"document_id": "card_protocol_001",
"document_type": "treatment_protocol",
"department": "cardiology",
"date": "2025-01-10",
"author_role": "department_head",
"text": """
Cardiology Protocol — Heart Failure with Reduced EF (HFrEF)
First-line therapy:
- ACE inhibitor: ramipril 2.5–10 mg daily (or ARB if ACE-intolerant).
- Beta-blocker: bisoprolol 1.25–10 mg daily, carvedilol 3.125–25 mg BID,
or metoprolol succinate 12.5–200 mg daily. Titrate every 2 weeks.
- MRA: spironolactone 25–50 mg daily for NYHA class II–IV
if eGFR > 30 and K+ < 5.0.
Target: Symptomatic improvement. Reassess LVEF at 3–6 months.
Device therapy (ICD/CRT) if LVEF ≤ 35% after 3 months optimal therapy.
""",
},
{
"document_id": "psych_note_001",
"document_type": "clinical_note",
"department": "psychiatry",
"date": "2025-03-18",
"author_role": "psychiatrist",
"text": """
Psychiatry intake note — [NAME REDACTED], [AGE REDACTED]-year-old.
Presenting with major depressive episode, PHQ-9 score 18 (severe).
No current suicidal ideation. Started sertraline 50 mg daily.
Psychotherapy referral placed. Follow-up in 2 weeks.
Safety plan documented. Family support confirmed present.
""",
},
{
"document_id": "onco_note_001",
"document_type": "clinical_note",
"department": "oncology",
"date": "2025-03-20",
"author_role": "oncologist",
"text": """
Oncology note — [NAME REDACTED].
Diagnosis: Stage IIIA non-small cell lung cancer, adenocarcinoma.
EGFR mutation positive (exon 19 deletion).
Plan: Osimertinib 80 mg daily (first-line EGFR-targeted therapy).
Baseline CT chest/abdomen/pelvis completed. Brain MRI negative.
Next imaging review in 8 weeks. Antiemetics PRN, skin care for rash.
""",
},
]
# ── Step 1: De-identification ─────────────────────────────────────────────────
# For production use Presidio:
# from presidio_analyzer import AnalyzerEngine
# from presidio_anonymizer import AnonymizerEngine
# analyzer, anonymizer = AnalyzerEngine(), AnonymizerEngine()
# result = analyzer.analyze(text=raw, entities=[...], language="en")
# clean = anonymizer.anonymize(text=raw, analyzer_results=result).text
#
# This demo applies lightweight regex to already-synthetic notes.
_HIPAA_PATTERNS = [
(r"\b\d{3}-\d{2}-\d{4}\b", "[SSN]"), # SSN
(r"\bMRN[-:\s]*\d{4,10}\b", "[MRN]"), # medical record #
(r"\b\d{1,2}/\d{1,2}/\d{2,4}\b", "[DATE]"), # dates
(r"\b[A-Z][a-z]+ [A-Z][a-z]+\b", "[NAME]"), # names (simple)
(r"\b\d{3}[-.\s]\d{3}[-.\s]\d{4}\b", "[PHONE]"), # phone
(r"\b[a-zA-Z0-9._%+-]+@[a-zA-Z0-9.-]+\.\w+\b","[EMAIL]"), # email
(r"\b\d{5}(?:-\d{4})?\b", "[ZIP]"), # zip
(r"\b(?:https?://)\S+", "[URL]"), # URLs
(r"\b\d{1,3}\.\d{1,3}\.\d{1,3}\.\d{1,3}\b", "[IP]"), # IP addresses
]
def deidentify(text: str) -> str:
"""Remove HIPAA Safe Harbor identifiers from text."""
for pattern, replacement in _HIPAA_PATTERNS:
text = re.sub(pattern, replacement, text)
return text.strip()
# ── Step 2: Chunking ──────────────────────────────────────────────────────────
def chunk(text: str, size: int = CHUNK_TOKENS, overlap: int = OVERLAP_TOKENS) -> list[str]:
"""Split text into overlapping token windows (whitespace tokenisation)."""
tokens = text.split()
chunks, start = [], 0
while start < len(tokens):
end = min(start + size, len(tokens))
chunks.append(" ".join(tokens[start:end]))
if end == len(tokens):
break
start += size - overlap
return chunks
# ── Step 3: Embedding ─────────────────────────────────────────────────────────
print(f"Loading embedding model: {EMBED_MODEL}")
_model = SentenceTransformer(EMBED_MODEL)
def embed(texts: list[str]) -> list[list[float]]:
return _model.encode(texts, normalize_embeddings=True).tolist()
# ── Step 4: Ingest into VectorAI DB ──────────────────────────────────────────
def _chunk_id(doc_id: str, idx: int) -> int:
"""Stable integer ID from (document_id, chunk_index)."""
h = hashlib.sha256(f"{doc_id}:{idx}".encode()).hexdigest()
return int(h[:15], 16)
def ingest(notes: list[dict]) -> None:
with VectorAIClient(VECTORAI_HOST) as client:
# Health check
info = client.health_check()
print(f"VectorAI DB connected version={info['version']}")
# Create collection (skip if already exists)
try:
client.collections.create(
name=COLLECTION,
vectors_config=VectorParams(size=VECTOR_DIM, distance=Distance.Cosine),
)
print(f"Collection '{COLLECTION}' created dim={VECTOR_DIM}")
except Exception as e:
if "exists" in str(e).lower():
print(f"Collection '{COLLECTION}' already exists — skipping create")
else:
raise
total_chunks = 0
for note in notes:
# De-identify FIRST — before chunking or embedding
clean_text = deidentify(note["text"])
# Chunk second
chunks = chunk(clean_text)
# Embed third
vectors = embed(chunks)
# Build PointStruct records with strict metadata schema
# All four metadata fields are REQUIRED — no optional fields.
points = [
PointStruct(
id=_chunk_id(note["document_id"], i),
vector=vectors[i],
payload={
# ── strict schema ──────────────────────────────────────
"document_type": note["document_type"], # required
"department": note["department"], # required — RBAC filter key
"date": note["date"], # required
"author_role": note["author_role"], # required
# ── retrieval helpers ──────────────────────────────────
"document_id": note["document_id"],
"chunk_index": i,
"text": chunks[i], # de-identified chunk text
},
)
for i in range(len(chunks))
]
client.points.upsert(COLLECTION, points)
total_chunks += len(chunks)
print(f" ✓ {note['document_id']} dept={note['department']} chunks={len(chunks)}")
print(f"\nIngestion complete — {len(notes)} documents, {total_chunks} chunks total")
if __name__ == "__main__":
ingest(RAW_NOTES)
This file performs the following actions:
- De-identifies data: The system removes all HIPAA identifiers from raw text before processing. Names, dates, and other sensitive fields are replaced with placeholders to prevent PHI from entering the system pipeline.
- Chunks the texts: The system splits the cleaned text into 512-token segments with a 50-token overlap. This overlap preserves context across boundaries, enhancing retrieval accuracy.
- Embeds the chunks: The model converts each chunk into a numerical vector using a local sentence-transformers model. This process captures semantic meaning while keeping all processing within the network.
- Stores with metadata: The system writes each chunk and its vector to VectorAI DB, along with necessary fields like document_type, department, date, and author_role. These fields support strict access control during queries.
Run the script by:
uv run ingest.py
You should see the following results after running the command:
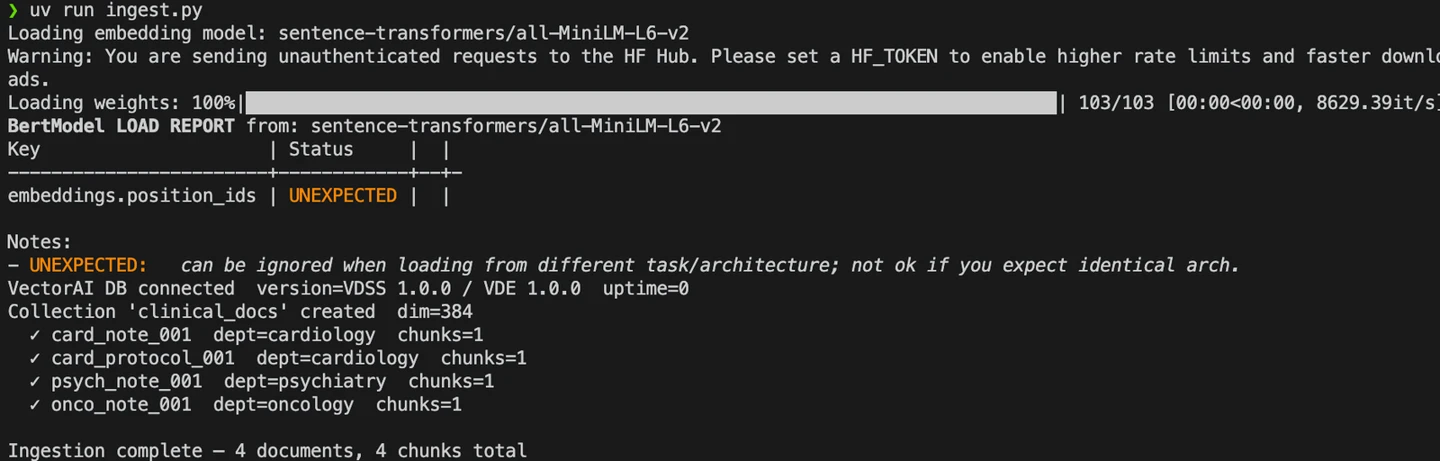
ingest.py execution
From the logs, you see that the ingestion pipeline writes chunks to VectorAI DB.
Step 3: Run your queries
Execute queries against your local RAG system and validate retrieval, access control, and audit logging.
Create a file query.py with the following contents:
import json
import datetime
import urllib.request
import urllib.error
from pathlib import Path
from actian_vectorai import VectorAIClient
from actian_vectorai import FilterBuilder, Field
from sentence_transformers import SentenceTransformer
# ── Config ─────────────────────────────────────────────────────────────────────
VECTORAI_HOST = "localhost:50052"
COLLECTION = "clinical_docs"
EMBED_MODEL = "sentence-transformers/all-MiniLM-L6-v2"
AUDIT_LOG = Path("./audit_logs/queries.jsonl") # volume-mounted path
# Ollama settings — set OLLAMA_ENABLED=True once `ollama serve` is running
OLLAMA_ENABLED = False # flip to True when Ollama is ready
OLLAMA_URL = "https://localhost:11434/api/generate"
OLLAMA_MODEL = "mistral" # or "llama3.2:3b" for lower hardware
# ── RBAC: role → allowed departments ──────────────────────────────────────────
# Access is enforced as a MUST filter at the database level.
# A scheduling_bot cannot reach clinical notes; cardiology cannot see psychiatry.
ROLE_PERMISSIONS = {
"cardiology_clinician": ["cardiology"],
"oncology_clinician": ["oncology"],
"general_practitioner": ["cardiology", "oncology", "general"],
"admin": ["cardiology", "oncology", "psychiatry", "general"],
"scheduling_bot": ["scheduling"], # no clinical note access
}
class AccessDeniedError(Exception):
pass
def allowed_departments(role: str) -> list[str]:
if role not in ROLE_PERMISSIONS:
raise AccessDeniedError(f"Unknown role '{role}' — access denied by default.")
return ROLE_PERMISSIONS[role]
# ── Embedding (reuse the same model as ingest.py) ─────────────────────────────
print(f"Loading embedding model: {EMBED_MODEL}")
_model = SentenceTransformer(EMBED_MODEL)
def embed(text: str) -> list[float]:
return _model.encode([text], normalize_embeddings=True).tolist()[0]
# ── Step 5: Search with department MUST filter ─────────────────────────────────
def retrieve(query_vec: list[float], departments: list[str], top_k: int = 5) -> list[dict]:
"""
Hybrid retrieval: vector similarity + metadata MUST filter.
Results from departments outside the allowed list are impossible —
the filter is applied at the database level, not in application code.
"""
results = []
with VectorAIClient(VECTORAI_HOST) as client:
for dept in departments:
hits = client.points.search(
collection_name=COLLECTION,
vector=query_vec,
limit=top_k,
# MUST filter — department equality enforced at DB level
filter=FilterBuilder().must(Field("department").eq(dept)).build(),
)
for hit in hits:
payload = getattr(hit, "payload", {}) or {}
results.append({
"score": round(getattr(hit, "score", 0.0), 4),
"document_id": payload.get("document_id"),
"document_type": payload.get("document_type"),
"department": payload.get("department"),
"date": payload.get("date"),
"author_role": payload.get("author_role"),
"chunk_index": payload.get("chunk_index"),
"text": payload.get("text", ""),
})
results.sort(key=lambda r: r["score"], reverse=True)
return results[:top_k]
# ── Step 6: LLM answer via Ollama ─────────────────────────────────────────────
_RAG_SYSTEM = (
"You are a clinical decision support assistant. "
"Answer ONLY using the context passages below. "
"Do NOT use external knowledge or make assumptions. "
"Cite each fact as [Doc N]. "
"If the context is insufficient, say: 'I cannot answer from the available documents.'"
)
def build_context(chunks: list[dict]) -> str:
return "\n\n".join(
f"[Doc {i+1}] ({c['document_type']}, dept={c['department']}, "
f"date={c['date']}, role={c['author_role']})\n{c['text']}"
for i, c in enumerate(chunks)
)
def generate(query_text: str, chunks: list[dict]) -> str:
if not OLLAMA_ENABLED:
# Return raw retrieved context when LLM is disabled
return "[LLM disabled — set OLLAMA_ENABLED=True]\n\n" + build_context(chunks)
context = build_context(chunks)
prompt = f"{_RAG_SYSTEM}\n\nContext:\n{context}\n\nQuestion: {query_text}\n\nAnswer:"
payload = json.dumps({
"model": OLLAMA_MODEL,
"prompt": prompt,
"stream": False,
"options": {"num_predict": 400},
}).encode()
try:
req = urllib.request.Request(
OLLAMA_URL,
data=payload,
headers={"Content-Type": "application/json"},
method="POST",
)
with urllib.request.urlopen(req, timeout=30) as resp:
data = json.loads(resp.read())
return data.get("response", "").strip()
except urllib.error.URLError as e:
return f"[Ollama unreachable: {e}]\n\nRetrieved context:\n{build_context(chunks)}"
def write_audit(record: dict) -> None:
AUDIT_LOG.parent.mkdir(parents=True, exist_ok=True)
with open(AUDIT_LOG, "a", encoding="utf-8") as f:
f.write(json.dumps(record, ensure_ascii=False) + "\n")
# ── Public query entry point ───────────────────────────────────────────────────
def query(user_id: str, role: str, query_text: str, top_k: int = 5) -> dict:
"""
Execute a role-gated RAG query.
Returns:
{answer, retrieved_docs, access_denied, error}
"""
timestamp = datetime.datetime.now(datetime.timezone.utc).isoformat()
# RBAC check — before anything else
try:
departments = allowed_departments(role)
except AccessDeniedError as e:
write_audit({
"timestamp": timestamp, "user_id": user_id, "role": role,
"department": "DENIED", "query_text": query_text,
"retrieved_docs": [], "answer_provided": False, "access_denied": True,
"denial_reason": str(e),
})
return {"answer": f"Access denied: {e}", "retrieved_docs": [], "access_denied": True}
# Embed → retrieve (with MUST filter) → generate
q_vec = embed(query_text)
chunks = retrieve(q_vec, departments, top_k)
answer = generate(query_text, chunks)
doc_refs = [
{"document_id": c["document_id"], "chunk_index": c["chunk_index"],
"department": c["department"], "document_type": c["document_type"],
"score": c["score"]}
for c in chunks
]
# Audit log — every query, regardless of outcome
write_audit({
"timestamp": timestamp,
"user_id": user_id,
"role": role,
"department": ",".join(departments),
"query_text": query_text,
"retrieved_docs": doc_refs,
"answer_provided": True,
"access_denied": False,
})
return {"answer": answer, "retrieved_docs": doc_refs, "access_denied": False}
# ── Demo runs ─────────────────────────────────────────────────────────────────
if __name__ == "__main__":
separator = "─" * 60
# ── Query 1: authorised cardiology query ────────────────────────────────
print(f"\n{separator}")
print("QUERY 1 — cardiology_clinician (authorised)")
print(separator)
r1 = query(
user_id="dr_chen_007",
role="cardiology_clinician",
query_text="What beta-blocker is recommended for heart failure with reduced ejection fraction?",
)
print(f"\nAnswer:\n{r1['answer']}\n")
print("Retrieved sources:")
for d in r1["retrieved_docs"]:
print(f" score={d['score']} [{d['document_type']}] dept={d['department']} "
f"doc={d['document_id']} chunk={d['chunk_index']}")
# ── Query 2: scheduling bot tries to access clinical notes ───────────────
print(f"\n{separator}")
print("QUERY 2 — scheduling_bot (attempting clinical note access)")
print(separator)
r2 = query(
user_id="bot_sched_01",
role="scheduling_bot",
query_text="What are the diagnosis notes for cardiology patients?",
)
if r2["access_denied"]:
print(f"\n✗ Access denied (as expected): {r2['answer']}")
else:
print(f"\nAnswer:\n{r2['answer']}")
print("Sources:", r2["retrieved_docs"])
# ── Query 3: cardiology query that must NOT return psychiatry notes ───────
print(f"\n{separator}")
print("QUERY 3 — cardiology_clinician (RBAC must exclude psychiatry)")
print(separator)
r3 = query(
user_id="dr_chen_007",
role="cardiology_clinician",
query_text="antidepressant dosing and patient management",
)
departments_returned = {d["department"] for d in r3["retrieved_docs"]}
cross_leak = "psychiatry" in departments_returned
print(f"\nDepartments in results: {departments_returned or 'none'}")
print(f"Cross-department leak: {'✗ LEAK DETECTED' if cross_leak else '✓ none — RBAC working correctly'}")
# ── Show audit log tail ───────────────────────────────────────────────────
print(f"\n{separator}")
print("AUDIT LOG → {AUDIT_LOG}")
print(separator)
if AUDIT_LOG.exists():
lines = AUDIT_LOG.read_text().strip().splitlines()
for line in lines[-3:]: # show last 3 entries
entry = json.loads(line)
print(json.dumps({
"timestamp": entry["timestamp"],
"user_id": entry["user_id"],
"role": entry["role"],
"query_text": entry["query_text"][:60] + "…",
"docs_accessed": len(entry["retrieved_docs"]),
"access_denied": entry["access_denied"],
}, indent=2))
else:
print("No audit log found — run ingest.py first.")
The script performs three core operations in a single flow.
- Enforces access control: The system checks the user’s role before retrieving any data. Each role is mapped to specific departments and enforced as a mandatory filter at the database level. The authorization layer immediately blocks and logs unauthorized roles.
- Retrieve and generate answers: The system embeds the query and retrieves relevant document chunks using vector search, with strict department filters applied. The results are then passed to a local LLM. If the LLM is disabled, the retrieved context is returned directly.
- Write audit logs: The system logs every query locally, including the user ID, role, query text, accessed documents, and access status. This creates a complete audit trail for compliance and review.
Run the script by:
uv run query.py
You should see the following results after running the command:
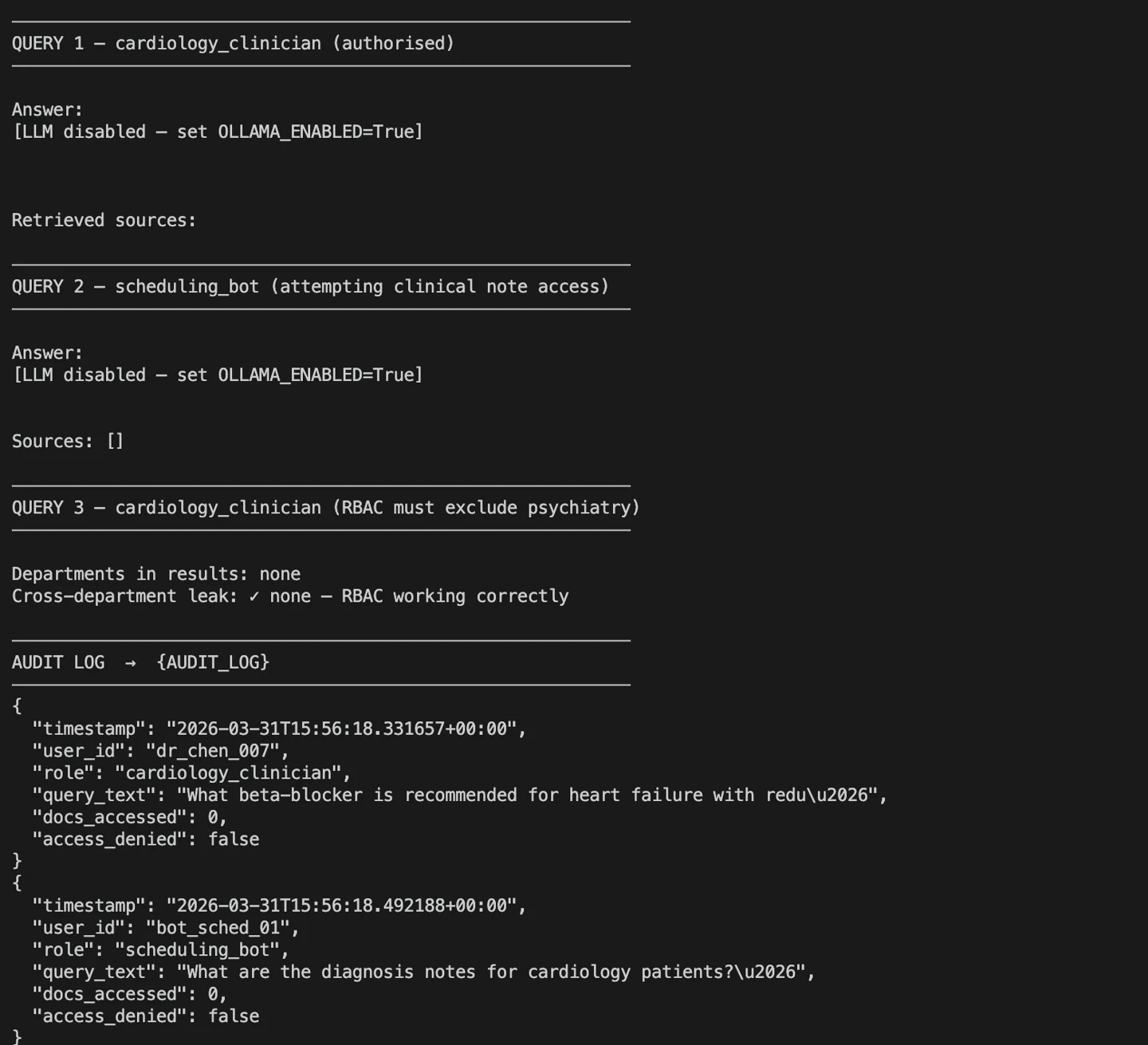
query.py execution
The output shows three test cases: one showing a valid clinician query, a denied access attempt, and a check for cross-department leakage. These confirm that RBAC and audit logging work correctly before you move to production.
Step 4: Configure the audit log
Store every query locally by using the volume mapping defined during deployment.
The Docker configuration mounts ./audit_logs from your host into the container. When you run queries, this creates a local folder named audit_logs with a file queries.jsonl.
The file contains the following entries:
{"timestamp": "2026-03-31T15:56:02.552088+00:00", "user_id": "dr_chen_007", "role": "cardiology_clinician", "department": "cardiology", "query_text": "What beta-blocker is recommended for heart failure with reduced ejection fraction?", "retrieved_docs": [], "answer_provided": true, "access_denied": false}
{"timestamp": "2026-03-31T15:56:03.098346+00:00", "user_id": "bot_sched_01", "role": "scheduling_bot", "department": "scheduling", "query_text": "What are the diagnosis notes for cardiology patients?", "retrieved_docs": [], "answer_provided": true, "access_denied": false}
{"timestamp": "2026-03-31T15:56:03.663767+00:00", "user_id": "dr_chen_007", "role": "cardiology_clinician", "department": "cardiology", "query_text": "antidepressant dosing and patient management", "retrieved_docs": [], "answer_provided": true, "access_denied": false}
{"timestamp": "2026-03-31T15:56:18.331657+00:00", "user_id": "dr_chen_007", "role": "cardiology_clinician", "department": "cardiology", "query_text": "What beta-blocker is recommended for heart failure with reduced ejection fraction?", "retrieved_docs": [], "answer_provided": true, "access_denied": false}
{"timestamp": "2026-03-31T15:56:18.492188+00:00", "user_id": "bot_sched_01", "role": "scheduling_bot", "department": "scheduling", "query_text": "What are the diagnosis notes for cardiology patients?", "retrieved_docs": [], "answer_provided": true, "access_denied": false}
{"timestamp": "2026-03-31T15:56:18.569824+00:00", "user_id": "dr_chen_007", "role": "cardiology_clinician", "department": "cardiology", "query_text": "antidepressant dosing and patient management", "retrieved_docs": [], "answer_provided": true, "access_denied": false}Each line represents a single query event. The log captures who made the request, their role, the department scope, the query text, and whether access was allowed. This file lives entirely on your infrastructure and gives you a complete, verifiable audit trail for every interaction with PHI.
Wrapping Up
A BAA was never enough because it does not control how your application handles PHI. You solved that by keeping all data, queries, and logs inside your network.
You now have a RAG system that enforces role-based access, retrieves only authorized data, and logs every interaction locally, without external APIs or third-party exposure.
Apply this pattern to other regulated systems. Refer to the VectorAI DB documentation and GitHub repository for updates and implementation details.
Join the community and learn more about Actian.





